Many vital processes with relation to biological phenomena such as heart beating, blood flowing and neural activities are often highly dynamic, changing in a period as short as milliseconds within the scope of 3D space. Fast and high-resolution microscopic observation of these dynamic processes is a very significant but challenging task, developments of which will render a great push to the research into the fields of neurosciences, cardiology, cytobiology and the like. For all the currently existing mainstream optical microscopic imaging technologies including confocal, two-photon and light-sheet microscopy, it is necessary to acquire multiple frames of planar images by A-scanning for the purpose of reconstructing 3D signals, so the rate of volumetric imaging is generally limited to seconds to minutes, which makes it hard to capture dynamic biological processes.
In response to the problem above, a research group led by Fei Peng from the School of Optical and Electronic Information published a long research article titled “Real-time volumetric reconstruction of biological dynamics with light-field microscopy and deep learning” in the international journal of Nature Methods in conjunction with Gao Shangbang-led Research Group from the College of Life Sciences & Technology, HUST, and TzungHsiai-led Research Group from UCLA on February 11, proposing a deep learning-based light-field imaging method and realizing the 3D observation of dynamic processes featuring a long period, high speed and high resolution.

The light-field 3D microscopy requiring no scanning is applied in this research to record the dynamic processes of samples at a frame rate of 100 HZ, and, during the research, an information conversion neural network of deep learning-based view-channel-depth (VCD) was first created, which can restore the 3D distribution of time-dependent signal changes from the recorded 2D light-field image sequence to reconstruct 3D fast dynamic processes in real time. With this method, the problem is solved that consideration cannot be given to both speediness and accuracy due to the space-bandwidth product (optical throughout) restricting the current 3D imaging technology, thus the milliseconds-long 3D dynamic biological processes of living samples can be successfully captured at single cell resolution, and 2D-3D images can be reconstructed at a speed of as high as tens of volumes per seconds nearly in real time. Based on this technology, Fei Feng-led Research Group and Gao Shangbang-led Research Group worked together in observing the neural activities and behavior of a freely moving threadworm at a volumetric rate of 100 HZ. The video of reconstructed 3D images could definitely tell the strength of a single neuron and its change in the spatial position (Fig. 1a). Benefited from the accuracy of a single cell and the milliseconds-long imaging rate, researchers quantitatively revealed the relations between the fluctuation trend of motoneuron Ca signals of a threadworm and its behavior (speed and behavior pattern), providing an important reference for the neurobehavior of a threadworm. Fei Peng-led Research Group also conducted further cooperation with Hsiai-led Research Group to demonstrate the application of this technology in signalizing the imaging of denser heart beating and blood flowing of a living zebra fish (Fig. 1b), reconstructing the transient signals of myocardium beating and blood flowing at a volumetric rate of 200 HZ and completing the corresponding quantitative analysis in cardiac hemodynamics.
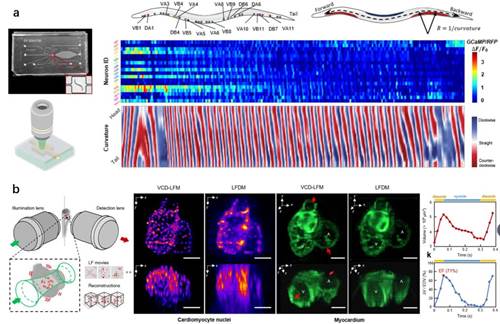
Fig.1: Deep learning-based light-field imaging captures the behavior of a freely moving threadworm, the myocardium beating of a zebra fish’s embryo, the flowing of haemocytes and other highly dynamic processes at the single cell level
The research contributes to the development of a new type of light-field microtechnique, with which milliseconds-long biological processes, hard to be clearly captured by currently existing microscopes, can be observed. This technology’s advantages in properties and application prospect in the research into life sciences have been fully shown by the fast and accurate imaging of the behavior of a freely moving threadworm and the heart beating and blood flowing of a zebra fish and by the quantitative biological analysis based on this.

Wang Zhaoqiang, a 2018 college graduate (currently studying at UCLA for his doctoral degree), Zhu Lanxin, a 2018 doctoral candidate, and Zhang Hao, a 2019 doctoral candidate, from the School of Optical and Electronic Information, HUST, are the co-first authors of the paper. Prof. Fei Peng from the School of Optical and Electronic Information, HUST, Prof. Gao Shangbang from the College of Life Science & Technology, HUST, and Prof. Hsiai from the David Geffen College of Medicine, UCLA, are the joint corresponding authors of the paper.
This research was conducted and accomplished with the financial aids from the Key R&D Program of the Ministry of Science and Technology, the General Program of the NFSC, the Key Instruments Development Project of the NFSC, the International Cooperation Key Project of the NFSC, the Innovation Funds of Wuhan National Laboratory for Optoelectronics (WNLO) and the Funds of the National Institute of Health (NIH)
Link to the paper: https://rdcu.be/cfaow
